2026

From Mind to Motion: An Interdisciplinary Review on NLP for the Impact of Online False Information on Beliefs and Behaviors
Mung Yao Jia , Yeaeun Gong, Dong Wang
Under review. 2026
From Mind to Motion: An Interdisciplinary Review on NLP for the Impact of Online False Information on Beliefs and Behaviors
Mung Yao Jia , Yeaeun Gong, Dong Wang
Under review. 2026
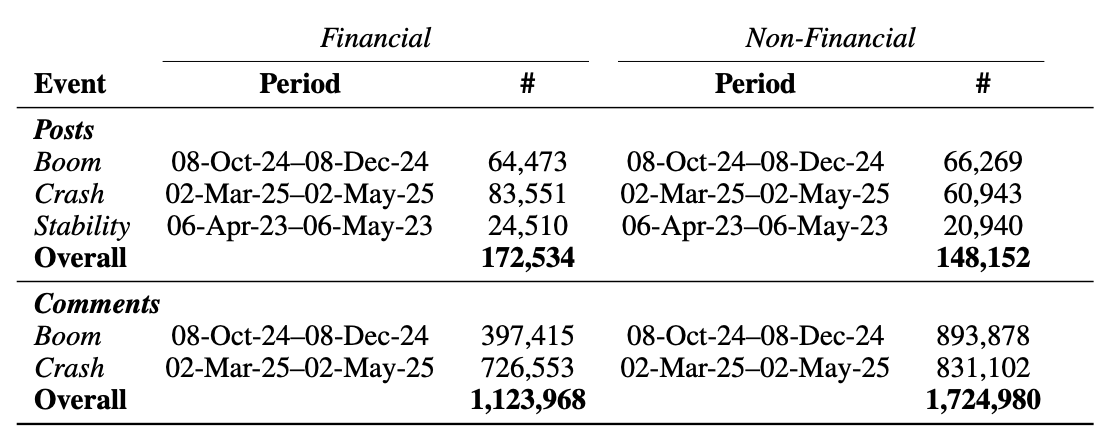
Are Gains Quiet and Losses Loud? Emotional Responses to Financial Booms and Crashes Online
Aryan Ramchandra Kapadia*, Niharika Bhattacharjee*, Mung Yao Jia* , Ishq Gupta, Dong Wang, Koustuv Saha (* equal contribution)
Digital Minds Workshop (DM) @ ICWSM 2026
Are Gains Quiet and Losses Loud? Emotional Responses to Financial Booms and Crashes Online
Aryan Ramchandra Kapadia*, Niharika Bhattacharjee*, Mung Yao Jia* , Ishq Gupta, Dong Wang, Koustuv Saha (* equal contribution)
Digital Minds Workshop (DM) @ ICWSM 2026
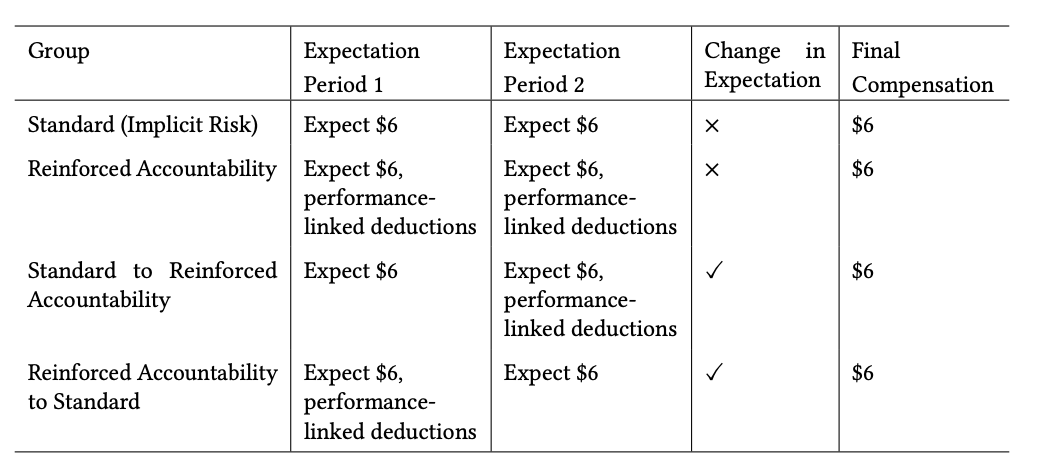
Dynamic Compensation Can Enhance User Engagement by Triggering Sensitivity to Financial Losses in Crowd-sourced Studies
Catalina Gomez*, Mung Yao Jia* , Sue Min Cho, Chien-Ming Huang, Mathias Unberath (* equal contribution)
Proceedings of the 2026 CHI Conference on Human Factors in Computing Systems 2026 Honourable Mention Award
Participation in crowd-sourced user studies is often driven by monetary incentives. However, standard payment schemes that reward completion unless responses are of poor quality may not invoke sufficient accountability. By compromising user engagement, a lack of accountability can affect data quality and the study’s ecological validity. Here, we investigate alternative compensation strategies that manipulate payment framing and evaluate their impact on engagement through task effort, outcomes, and perception. We compared a standard scheme with implicit rejection risk to a reinforced accountability condition with explicit performance-linked deductions, and two dynamic conditions that unexpectedly switched strategies. In a study with 106 Prolific participants on an image captioning task, we found that only shifting from implicit risk to reinforced accountability significantly increased engagement, likely due to loss aversion after participants had already invested time. The reverse shift decreased effort as observed in the standard group. Our results highlight the importance of carefully designing compensation schemes.
Dynamic Compensation Can Enhance User Engagement by Triggering Sensitivity to Financial Losses in Crowd-sourced Studies
Catalina Gomez*, Mung Yao Jia* , Sue Min Cho, Chien-Ming Huang, Mathias Unberath (* equal contribution)
Proceedings of the 2026 CHI Conference on Human Factors in Computing Systems 2026 Honourable Mention Award
Participation in crowd-sourced user studies is often driven by monetary incentives. However, standard payment schemes that reward completion unless responses are of poor quality may not invoke sufficient accountability. By compromising user engagement, a lack of accountability can affect data quality and the study’s ecological validity. Here, we investigate alternative compensation strategies that manipulate payment framing and evaluate their impact on engagement through task effort, outcomes, and perception. We compared a standard scheme with implicit rejection risk to a reinforced accountability condition with explicit performance-linked deductions, and two dynamic conditions that unexpectedly switched strategies. In a study with 106 Prolific participants on an image captioning task, we found that only shifting from implicit risk to reinforced accountability significantly increased engagement, likely due to loss aversion after participants had already invested time. The reverse shift decreased effort as observed in the standard group. Our results highlight the importance of carefully designing compensation schemes.
2025
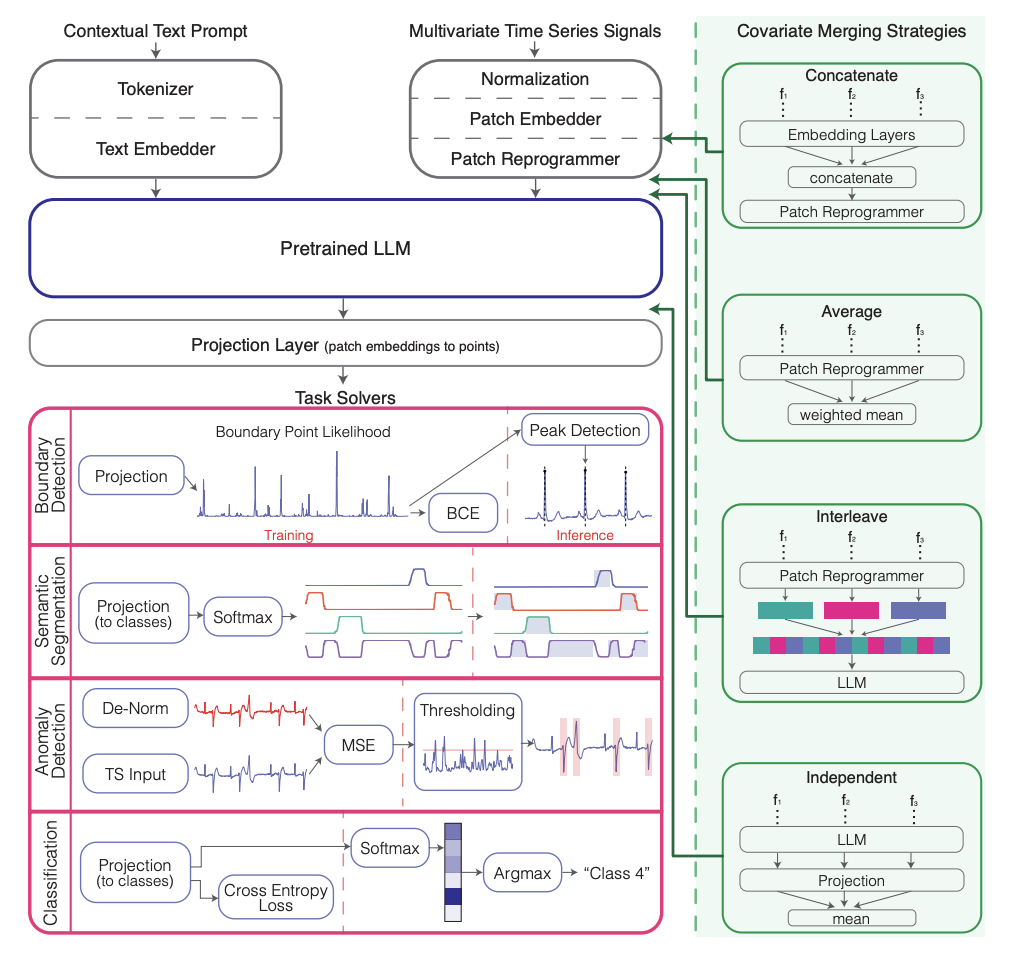
MedTsLLM: Medical Time Series Analysis Using Multimodal LLMs
Nimeesha Chan, Felix Parker, Chi Zhang, William Bennett, Mung Yao Jia , James Fackler, Kimia Ghobadi
IEEE Journal of Biomedical and Health Informatics 2025
Traditional machine learning approaches for biomedical time series analysis face fundamental limitations when integrating the heterogeneous data types essential for comprehensive clinical understanding. Physiological signals must be interpreted within rich clinical contexts that include patient history, current medications, and treatment protocols-information typically stored as unstructured text that conventional time series models cannot effectively utilize. We propose MedTsLLM, a multimodal model that aims to address this critical gap by integrating numerical physiological signals with natural language clinical information through large language models (LLMs). Our framework incorporates patch reprogramming for time series-LLM alignment and introduces two key innovations: novel covariate handling strategies that capture complex physiological relationships, and contextual prompting mechanisms that incorporate patient-specific information. MedTsLLM addresses four clinically significant tasks within a unified architecture: semantic segmentation, boundary detection, anomaly detection, and classification. Through comprehensive evaluation across diverse medical domains, including ECG analysis, respiratory monitoring, and cardiac arrhythmia detection, our approach consistently outperforms state-of-the-art baselines across all tasks and datasets. These results demonstrate the transformative potential of multimodal LLMs for biomedical signal analysis, enabling clinicians to extract deeper insights from physiological data while leveraging comprehensive clinical context to enhance diagnostic accuracy, patient monitoring, and personalized treatment decisions.
MedTsLLM: Medical Time Series Analysis Using Multimodal LLMs
Nimeesha Chan, Felix Parker, Chi Zhang, William Bennett, Mung Yao Jia , James Fackler, Kimia Ghobadi
IEEE Journal of Biomedical and Health Informatics 2025
Traditional machine learning approaches for biomedical time series analysis face fundamental limitations when integrating the heterogeneous data types essential for comprehensive clinical understanding. Physiological signals must be interpreted within rich clinical contexts that include patient history, current medications, and treatment protocols-information typically stored as unstructured text that conventional time series models cannot effectively utilize. We propose MedTsLLM, a multimodal model that aims to address this critical gap by integrating numerical physiological signals with natural language clinical information through large language models (LLMs). Our framework incorporates patch reprogramming for time series-LLM alignment and introduces two key innovations: novel covariate handling strategies that capture complex physiological relationships, and contextual prompting mechanisms that incorporate patient-specific information. MedTsLLM addresses four clinically significant tasks within a unified architecture: semantic segmentation, boundary detection, anomaly detection, and classification. Through comprehensive evaluation across diverse medical domains, including ECG analysis, respiratory monitoring, and cardiac arrhythmia detection, our approach consistently outperforms state-of-the-art baselines across all tasks and datasets. These results demonstrate the transformative potential of multimodal LLMs for biomedical signal analysis, enabling clinicians to extract deeper insights from physiological data while leveraging comprehensive clinical context to enhance diagnostic accuracy, patient monitoring, and personalized treatment decisions.
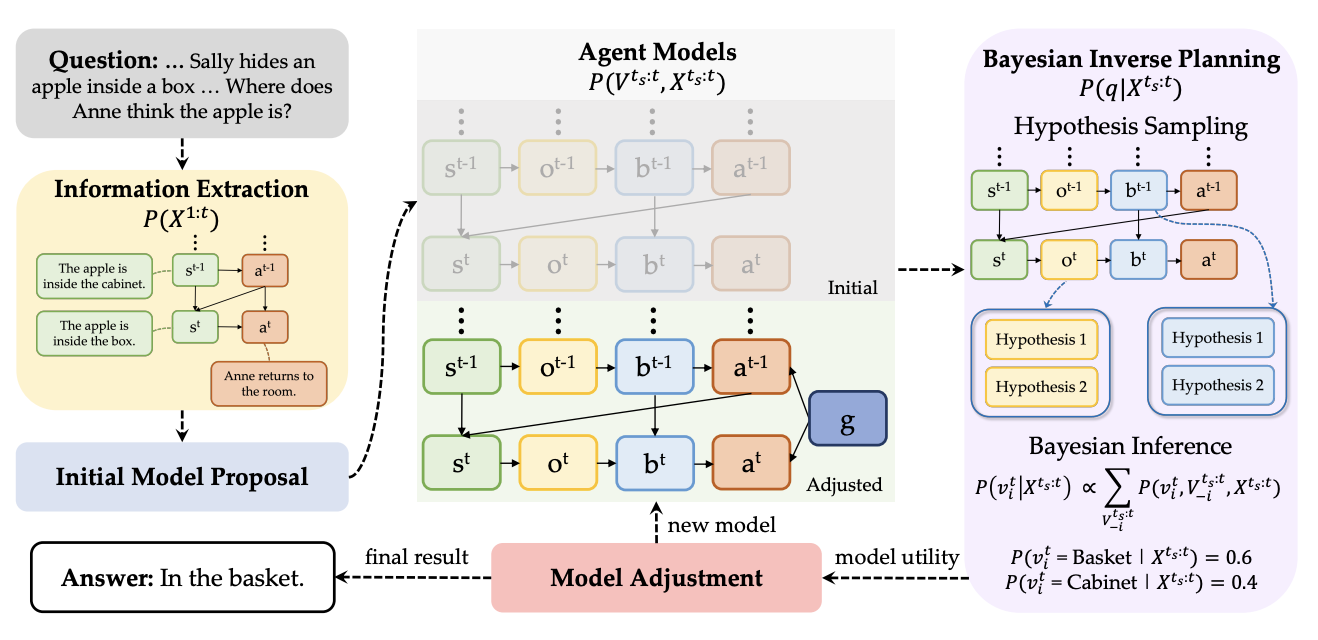
AutoToM: Scaling Model-based Mental Inference via Automated Agent Modeling
Zhining Zhang*, Chuanyang Jin*, Mung Yao Jia *, Shunchi Zhang*, Tianmin Shu (* equal contribution)
NeurIPS 2025 Spotlight
Theory of Mind (ToM), the ability to understand people’s minds based on their behavior, is key to developing socially intelligent agents. Current approaches to ToM reasoning either rely on prompting Large Language Models (LLMs), which are prone to systematic errors, or use handcrafted, rigid agent models for model-based inference, which are more robust but fail to generalize across domains. In this work, we introduce AutoToM, an automated agent modeling method for scalable, robust, and interpretable mental inference. Given a ToM problem, AutoToM first proposes an initial agent model and then performs automated Bayesian inverse planning based on this model, leveraging an LLM backend. Guided by inference uncertainty, it iteratively refines the model by introducing additional mental variables and/or incorporating more timesteps in the context. Across five diverse benchmarks, AutoToM outperforms existing ToM methods and even large reasoning models. Additionally, we show that AutoToM can produce human-like confidence estimates and enable online mental inference for embodied decision-making.
AutoToM: Scaling Model-based Mental Inference via Automated Agent Modeling
Zhining Zhang*, Chuanyang Jin*, Mung Yao Jia *, Shunchi Zhang*, Tianmin Shu (* equal contribution)
NeurIPS 2025 Spotlight
Theory of Mind (ToM), the ability to understand people’s minds based on their behavior, is key to developing socially intelligent agents. Current approaches to ToM reasoning either rely on prompting Large Language Models (LLMs), which are prone to systematic errors, or use handcrafted, rigid agent models for model-based inference, which are more robust but fail to generalize across domains. In this work, we introduce AutoToM, an automated agent modeling method for scalable, robust, and interpretable mental inference. Given a ToM problem, AutoToM first proposes an initial agent model and then performs automated Bayesian inverse planning based on this model, leveraging an LLM backend. Guided by inference uncertainty, it iteratively refines the model by introducing additional mental variables and/or incorporating more timesteps in the context. Across five diverse benchmarks, AutoToM outperforms existing ToM methods and even large reasoning models. Additionally, we show that AutoToM can produce human-like confidence estimates and enable online mental inference for embodied decision-making.
2024

MedTsLLM: Leveraging LLMs for Multimodal Medical Time Series Analysis
Nimeesha Chan, Felix Parker, William Bennett, Tianyi Wu, Mung Yao Jia , James Fackler, Kimia Ghobadi
Machine Learning for Healthcare Conference. 2024
The complexity and heterogeneity of data in many real-world applications pose significant challenges for traditional machine learning and signal processing techniques. For instance, in medicine, effective analysis of diverse physiological signals is crucial for patient monitoring and clinical decision-making and yet highly challenging. We introduce MedTsLLM, a general multimodal large language model (LLM) framework that effectively integrates time series data and rich contextual information in the form of text to analyze physiological signals, performing three tasks with clinical relevance: semantic segmentation, boundary detection, and anomaly detection in time series. These critical tasks enable deeper analysis of physiological signals and can provide actionable insights for clinicians. We utilize a reprogramming layer to align embeddings of time series patches with a pretrained LLM's embedding space and make effective use of raw time series, in conjunction with textual context. Given the multivariate nature of medical datasets, we develop methods to handle multiple covariates. We additionally tailor the text prompt to include patient-specific information. Our model outperforms state-of-the-art baselines, including deep learning models, other LLMs, and clinical methods across multiple medical domains, specifically electrocardiograms and respiratory waveforms. MedTsLLM presents a promising step towards harnessing the power of LLMs for medical time series analysis that can elevate data-driven tools for clinicians and improve patient outcomes.
MedTsLLM: Leveraging LLMs for Multimodal Medical Time Series Analysis
Nimeesha Chan, Felix Parker, William Bennett, Tianyi Wu, Mung Yao Jia , James Fackler, Kimia Ghobadi
Machine Learning for Healthcare Conference. 2024
The complexity and heterogeneity of data in many real-world applications pose significant challenges for traditional machine learning and signal processing techniques. For instance, in medicine, effective analysis of diverse physiological signals is crucial for patient monitoring and clinical decision-making and yet highly challenging. We introduce MedTsLLM, a general multimodal large language model (LLM) framework that effectively integrates time series data and rich contextual information in the form of text to analyze physiological signals, performing three tasks with clinical relevance: semantic segmentation, boundary detection, and anomaly detection in time series. These critical tasks enable deeper analysis of physiological signals and can provide actionable insights for clinicians. We utilize a reprogramming layer to align embeddings of time series patches with a pretrained LLM's embedding space and make effective use of raw time series, in conjunction with textual context. Given the multivariate nature of medical datasets, we develop methods to handle multiple covariates. We additionally tailor the text prompt to include patient-specific information. Our model outperforms state-of-the-art baselines, including deep learning models, other LLMs, and clinical methods across multiple medical domains, specifically electrocardiograms and respiratory waveforms. MedTsLLM presents a promising step towards harnessing the power of LLMs for medical time series analysis that can elevate data-driven tools for clinicians and improve patient outcomes.